Featured Applications
From glycoforms to biomarkers: assays that add specificity where it matters most
We apply platform-enabled glyco(proteo)mics to generate biomarker-ready molecular readouts from complex biological samples. Our work spans discovery, verification, and assay development, linking glycosylation and proteoform variation to biological mechanisms and clinically relevant phenotypes. Oncology is a key application area, and we also pursue disease settings where glycosylation reflects pathway activity, immune interactions, or tissue remodeling, making it a powerful layer for diagnostics and stratification.
A recurring theme is moving from abundance, only measurements to glycoform, and proteoform-resolved readouts, supported by targeted enrichment, high-sensitivity CE–MS, and reproducible analysis.
Featured application highlights
PSA: urinary PSA glycomics (PGA) and intact proteoform profiling, with extension toward serum workflows. Learn more
PAP: in-depth, site-specific and isomer-aware urinary PAP glycoproteomics. Learn more
CEA/CEACAM5: deep site-resolved mapping of a highly glycosylated cancer-associated protein. Learn more

Urinary PSA glycomics assay (PGA). Serum PSA concentration is used for early detection of prostate cancer, but clinical performance is limited by low specificity and sensitivity. Because PSA glycosylation has long been proposed as a more specific marker, we established a high-performance PSA Glycomics Assay (PGA) for urinary PSA that addresses a key bottleneck in the field: analytical sensitivity and specificity. After affinity purification from urine and tryptic digestion, PSA glycopeptides are analyzed by CE–ESI–MS, enabling in-depth relative quantitation of PSA glycoforms and, crucially, differentiation of α2,6- and α2,3-sialylated isomers, with α2,3-sialylation often discussed in the context of aggressive disease biology.
Prostate-Specific Antigen (PSA)
Extending towards serum PSA. While urine enables deep profiling with enrichment, clinically deployed PSA testing is serum-based. We are therefore translating and extending these workflows toward in-depth serum PSA analysis, supported by an Astellas ISR grant, with the goal of enabling glycoform- and proteoform-informed PSA readouts in a matrix and context that is directly relevant to clinical decision-making.
Related capability: Protein-of-interest enrichment + CE–ESI–MS glyco(proteo)mics, including isomer-aware sialylation profiling and intact proteoform analysis.
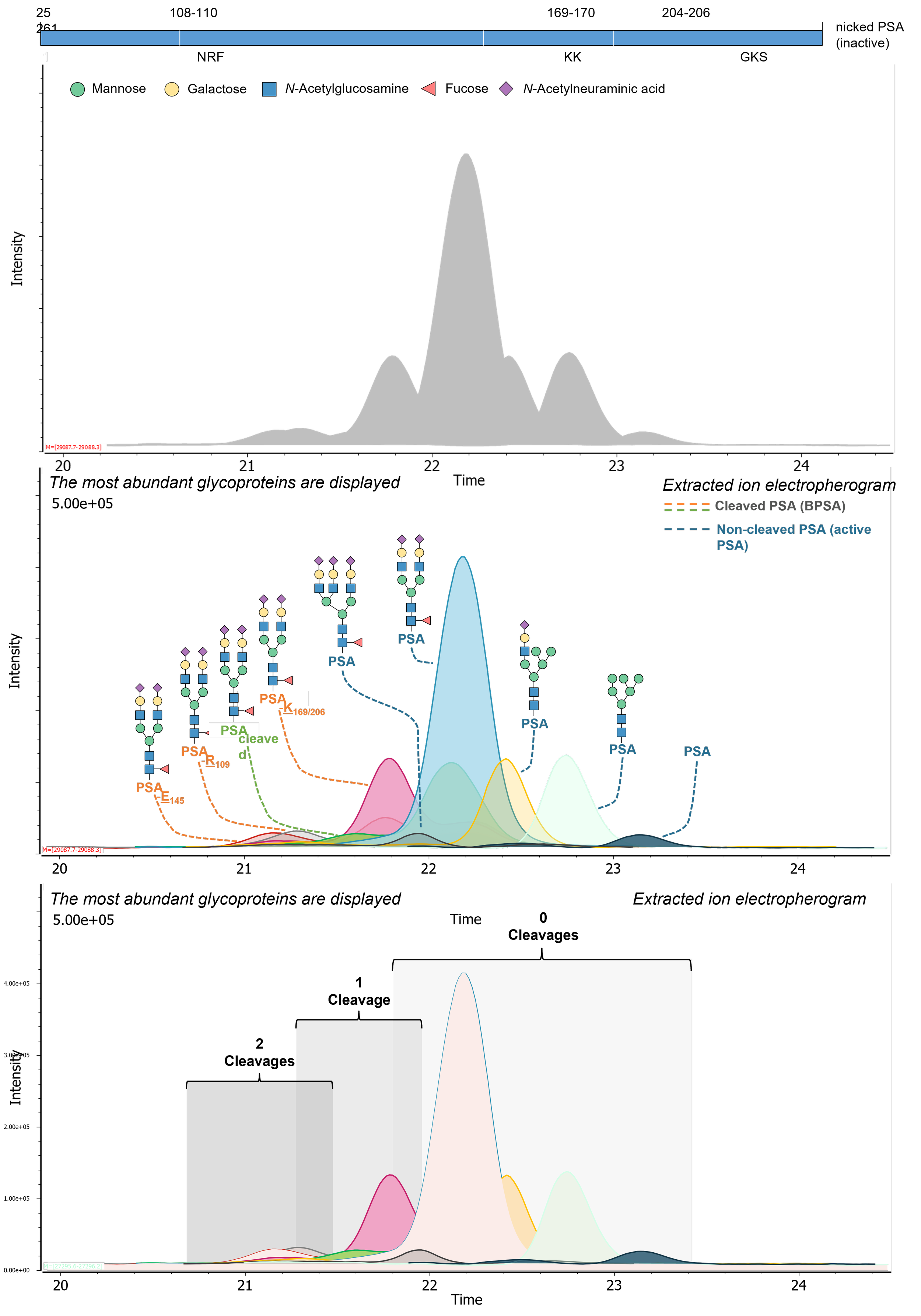
Complementary intact PSA proteoform profiling. Building on the glycopeptide-level workflow, we developed an intact protein CE–ESI–MS assay for affinity-purified urinary PSA to capture the PSA proteoform landscape with minimal additional sample treatment. This approach resolves proteolytic cleavage variants and provides a direct view on intact glycoforms (from non- to tri-sialylated complex-type glycans), including non-glycosylated PSA. By combining intact analysis with bottom-up glycopeptide profiling, we obtain orthogonal, complementary information, linking the contribution of cleaved vs non-cleaved PSA to the overall glycosylation profile and providing a stronger basis for future biomarker studies.
Why revisit PAP. PSA is the dominant clinical biomarker for prostate cancer, but serum PSA quantification has limited specificity and poor predictive value—driving continued efforts to find better tests. Before PSA, prostatic acid phosphatase (PAP) was used clinically, and when revisiting PAP specifically through the lens of glycosylation, it emerges as a promising biomarker candidate: prior studies suggest PAP glycoforms differ between PCa and non-PCa individuals, yet detailed characterization has been missing.
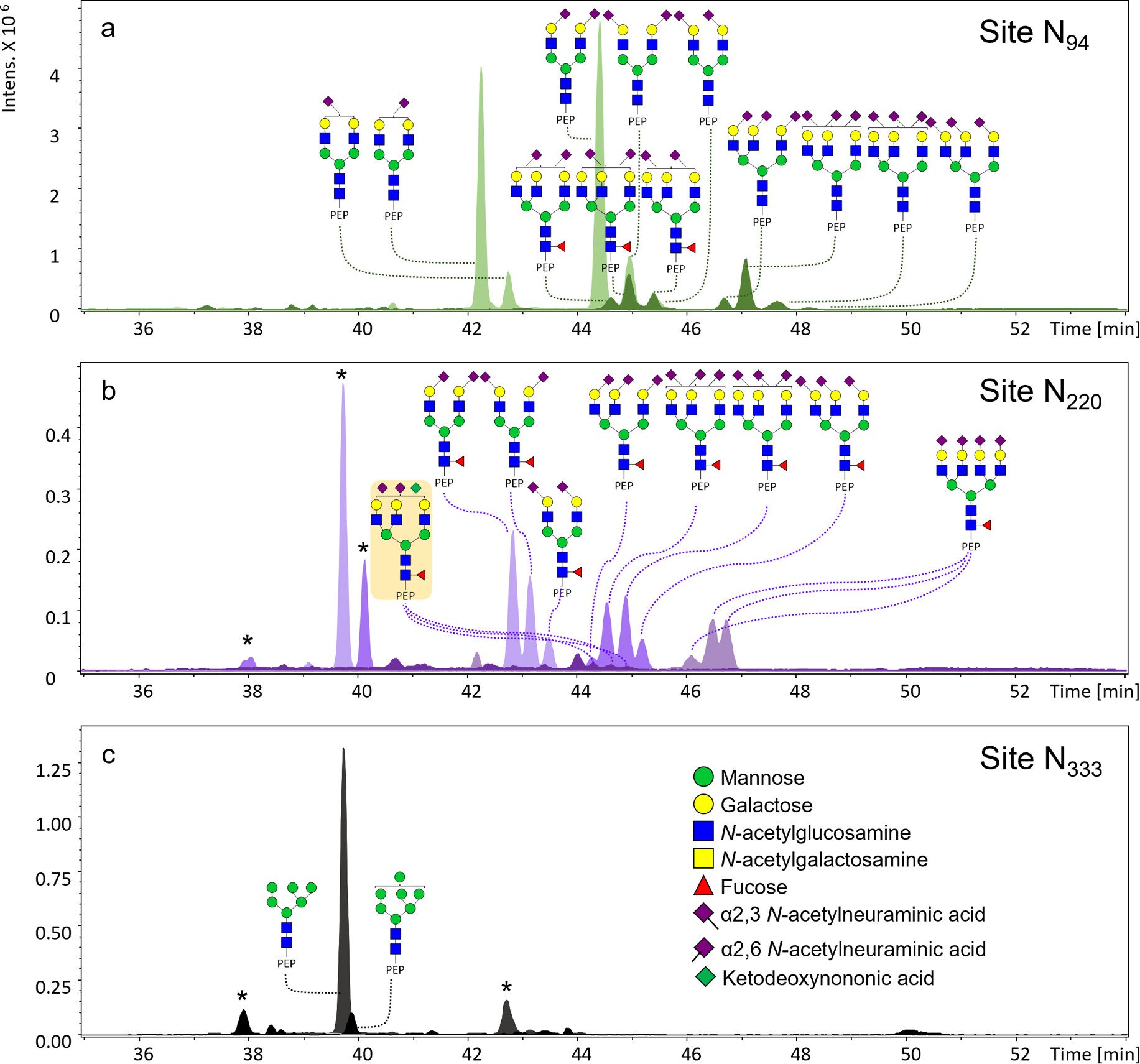
In-depth urinary PAP glycoproteomic assay. We established an in-depth glycoproteomic assay for urinary PAP to characterize both macroheterogeneity (which glycans are present) and microheterogeneity (how glycoforms vary at specific sites). After affinity purification from urine and proteolytic digestion, PAP was analyzed by capillary electrophoresis–mass spectrometry (CE–MS). Applying the workflow to a pooled digital rectal examination (DRE) urine sample, we characterized three N-glycosylation sites (N94, N220, N333) via N-glycopeptide analysis.
Isomer-aware site-resolved glycoprofiles for cohort studies. Taking sialic acid linkage isomers into account, we identified a rich site-specific glycan landscape, including 63, 27, and 4 N-glycan structures at the three sites, respectively. This assay provides a robust foundation for larger cohort studies aimed at determining whether PAP glycoforms can support improved early detection, prognosis, and stratification in prostate cancer.
Related capability: Protein-of-interest enrichment from urine + site-specific, isomer-aware glycoproteomics using CE–MS for deep glycoform profiling.
Prostatic Acid Phosphatase (PAP)
Carcinoembryonic Antigen (CEA)
Why CEA is a compelling (and challenging) target. Human carcinoembryonic antigen (CEA/CEACAM5) is an archetype of an extremely glycosylated tumor-associated protein: it has 28 potential N-glycosylation sites, and roughly ~60% of its molecular mass can be attributed to carbohydrates. CEA is frequently overexpressed and released by many solid tumors, including colorectal cancer, but despite its clinical relevance, site-specific glycosylation distributions had remained largely uncharted due to the extreme heterogeneity.
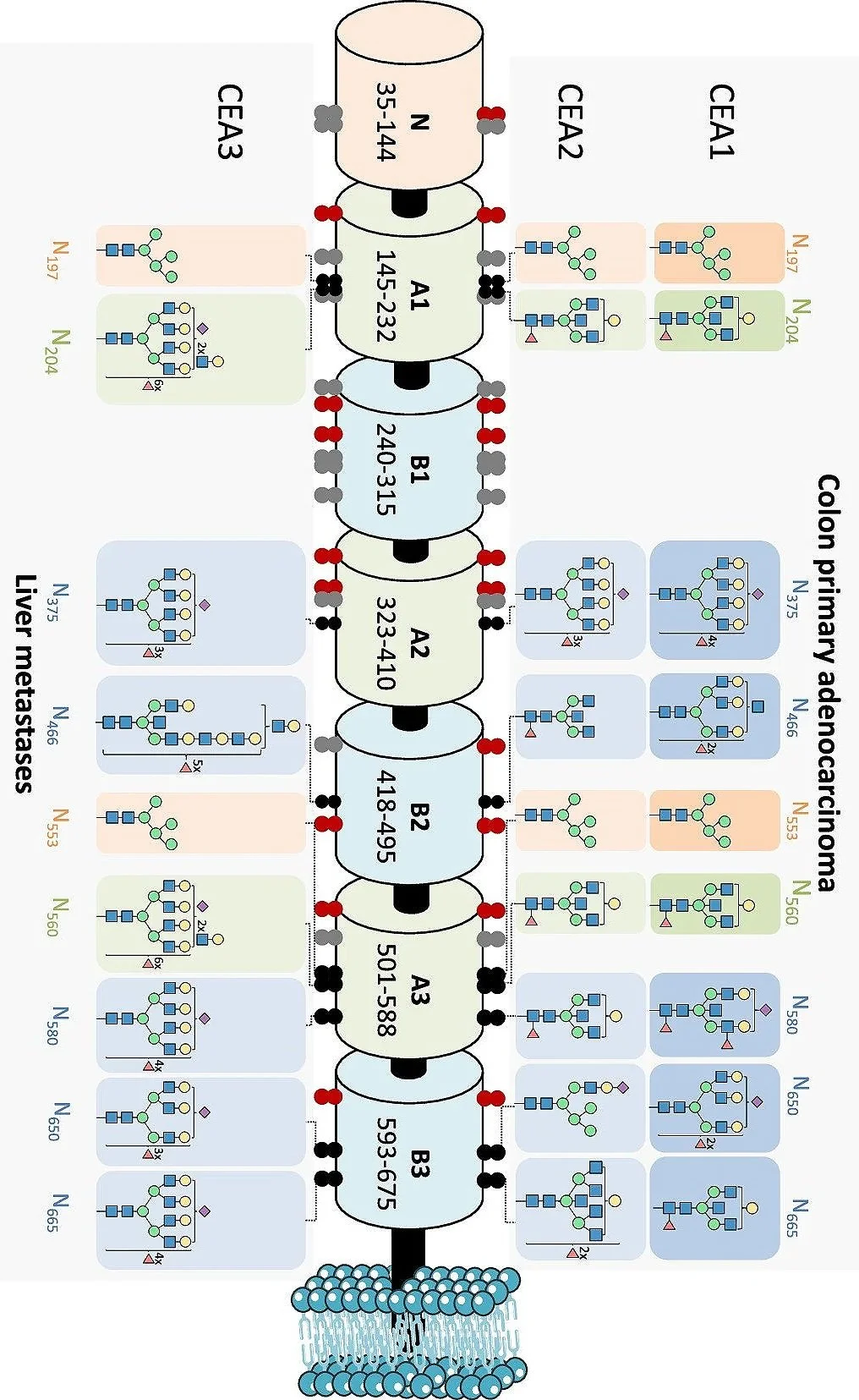
Deep site-resolved mapping enabled by sheathless CE–MS/MS. We investigated CEA purified from human colon carcinoma and human liver metastases and achieved high site coverage by combining a multienzymatic digestion strategy (trypsin, Glu-C, and Pronase) with sheathless CE–MS/MS. This enabled characterization of 21 out of 28 potential N-glycosylation sites with respect to occupancy and generated a rich molecular map, including 893 N-glycopeptides and 128 unique N-glycan compositions.
From heterogeneity to biomarker-relevant features. The results showed pronounced heterogeneity both within individual sites (microheterogeneity) and between sites (macroheterogeneity). Importantly, we observed site-specific differences between primary adenocarcinoma and metastatic tumor in features such as branching, bisection, sialylation, and fucosylation. These site-resolved, tumor-context-dependent glycosylation features provide a foundation for targeted follow-up aimed at improving diagnostics and monitoring of colorectal cancer progression and recurrence.
Related capability: Deep, site-resolved glycoproteomics of highly glycosylated proteins using tailored digestion strategies + sheathless CE–MS/MS, enabling occupancy and glycoform mapping across many sites.
Featured example: multi-glycan-class integration in AML (work performed at LUMC).
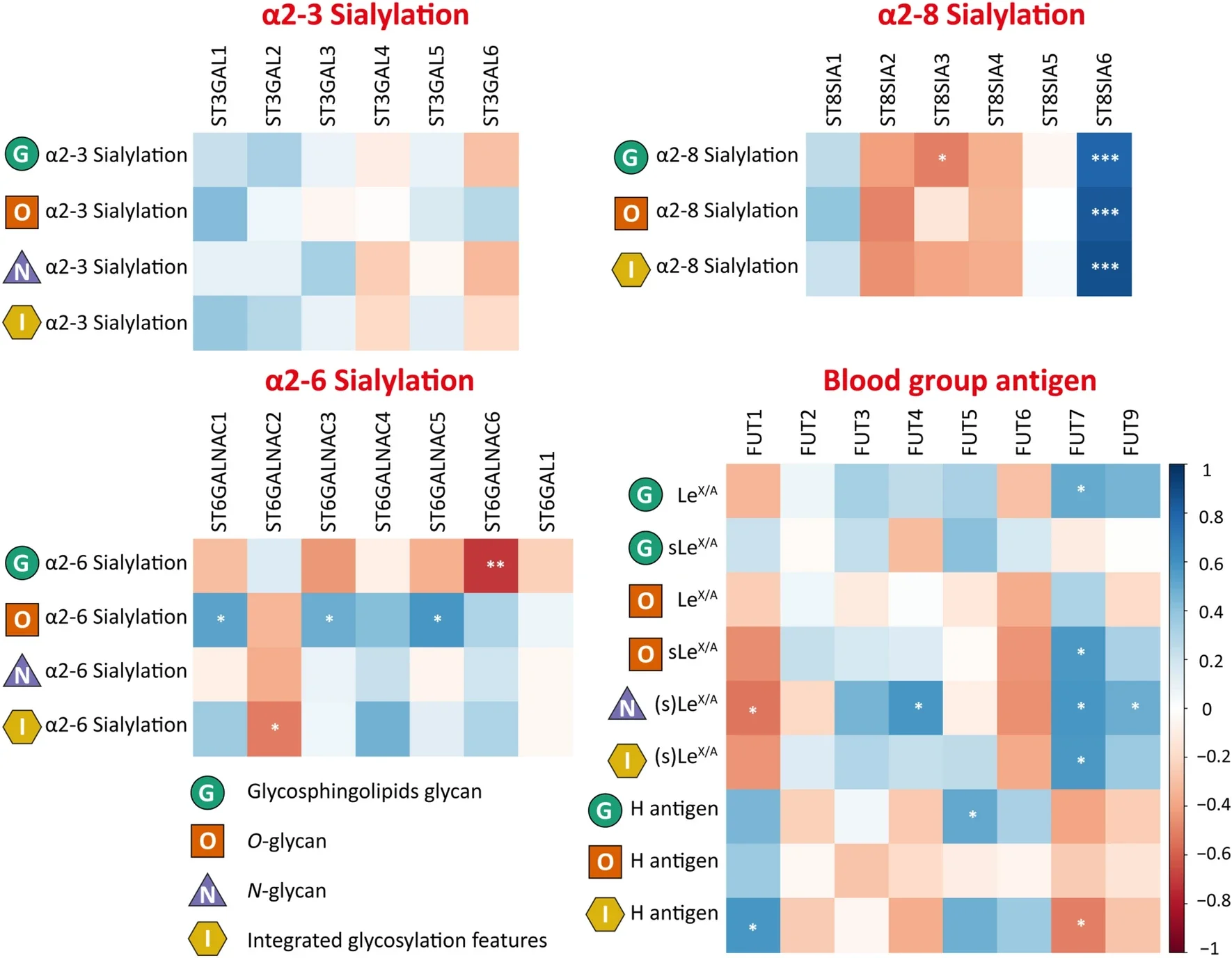
In a meta-omics study of acute myeloid leukemia (AML), we integrated N-glycomics, O-glycomics, and glycosphingolipid (GSL) glycomics from AML cell lines with transcriptomics from public datasets to link glycan phenotypes to glycosyltransferases and upstream regulatory networks. This approach highlighted FAB subtype-associated signatures (e.g., high (s)Le(x/a) in M5 and high sialylation in M6) and identified transcriptionally imprinted glycomic programs that were also reflected in primary AML datasets. This work was carried out at LUMC during my time there and illustrates the power of integrating multiple glycan classes with transcriptomics for mechanistic insight and stratification.
Integrated readouts across glycan classes
Biological questions rarely map to a single glycan class. Where it strengthens interpretation, we integrate multiple layers (such as N- and O-glycans, glycopeptides/glycoproteoforms, and glycolipids (glycosphingolipids)) to build a more complete molecular picture. This helps distinguish true biology from matrix or processing effects and supports clearer links to phenotype and mechanism.
Cross-validate findings across complementary glycan classes
Align glycosylation features with sample metadata and study design
Enable multi-layer biomarker hypotheses that can be followed up with targeted assays
Integrated readouts across glycan classes
Interested in collaboration or requesting measurements?