Analytical Platforms
Glycosylation in cells, samples, and disease, measured with exceptional sensitivity.
Our core expertise is building and applying high-sensitivity analytical platforms for glyco(proteo)mics. We combine sheathless capillary electrophoresis–mass spectrometry (CE–MS) with complementary MALDI-TOF-MS and MALDI-MSI strategies to extract molecular detail from complex biology, ranging from composition-level profiles to isomer-resolved and spatial glycosylation readouts.
CE–MS is the analytical backbone of the SUGAR Lab, enabling exceptionally sensitive glyco(proteo)mics through efficient separations and ultra-low-flow nano-ESI. We develop sheathless CE–MS workflows to characterize glycans, glycopeptides, and glycoproteoforms in complex biological matrices and limited material—supporting our drive toward single-cell-scale readouts.
Beyond abundance, biologically meaningful glycosylation changes often involve structural isomers—especially α2,6 vs α2,3 sialylation with different functional implications. We leverage the separation power of CE to resolve isomeric glycans and glycopeptides with minimal extra preparation, and we complement this with derivatization or neutral/zero-flow approaches to expand isomer distinction further.
MALDI–TOF–MS provides rapid, robust glycomics profiling and is particularly powerful for cohort-style studies and total plasma/serum N-glycome readouts. With very small input (as little as ~1 µL plasma/serum), we can generate profiles covering 50 up to 75 glycoforms, and, when paired with derivatization, add sialic-acid linkage specificity in a MALDI-compatible workflow.
To understand glycosylation in tissue context, we combine laser capture microdissection with sensitive downstream N- and O-glycomics to profile defined regions rather than bulk averages. In parallel, new MSI infrastructure at UMCG/FSE creates a strong foundation for expanding spatial omics capabilities, and developing MSI-based spatial glycomics is a clear near-future direction.
Together, these pillars let us move from sensitive detection to structural specificity and spatial context, turning glycosylation patterns into interpretable biology.

At the SUGAR Lab, CE–MS is more than an instrument coupling, it is the foundation of how we make glycosylation measurable when samples are complex and material is limited. We develop and use sheathless CE–MS (SCIEX) coupled to high-resolution mass spectrometry (e.g., timsTOF-SCP), enabling ultra-low nano-ESI flow rates (<10 nL/min) that push analytical sensitivity into regimes where subtle glycoforms become visible.
We build CE–MS workflows around efficient separations for charged and polar analytes, and we pair them with fit-for-purpose sample preparation (enrichment, labeling) and rigorous QC so that sensitivity is matched by interpretability. We also use CE–MS/MS to resolve structural glycan features and to map glycopeptide and glycoproteoform diversity—capturing heterogeneity that bulk measurements often obscure. This capability supports our broader aim of advancing glycomics toward single-cell-scale analysis.
What this platform enables
High sensitivity from limited material through ultra-low-flow ESI
High-efficiency separations that complement LC–MS for glycans and other polar/charged analytes
Interpretable glyco(proteo)mics workflows through targeted sample prep and QC
Our platform development is exemplified by our Nature Communications study demonstrating high-sensitivity CE–ESI–MS analysis of N-glycans using a sheathless interface.
Interested in using CE–MS to answer a biological question? Tell us your sample type and goal, we will propose a starting approach.
CE–MS (ultrasensitive glyco(proteo)mics
MALDI-TOF-MS for glyco(proteo)mics profiling
Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI–TOF–MS) provides a fast and information-rich readout of glycosylation, making it a powerful complement to separation-based approaches. In our work, MALDI–TOF–MS is particularly valuable for glycomics profiling (released glycans) and for studies where you want to compare many samples efficiently while still capturing biologically meaningful shifts in glycan patterns. Because MALDI measurements are rapid and robust, they can support cohort-style designs and exploratory screening, and it provides a high-throughput, robust complement to CE–MS, which we use for deeper, separation-resolved characterization.
A key strength is high-throughput total plasma/serum N-glycomics profiling: already ~1 µL of plasma or serum can be sufficient to generate a profile of 50 up to 75 glycoforms, including readouts that distinguish sialic-acid linkage variants when paired with the appropriate derivatization strategy. More broadly, MALDI is interesting for glyco(proteo)mics because glycans generate highly characteristic mass patterns, enabling sensitive detection of compositional shifts and, with the right chemistry, added structural specificity. For example, linkage-specific derivatization can encode α2,3 versus α2,6 sialylation into diagnostic mass differences, enabling fast discrimination in a MALDI-compatible workflow.
What this capability enables
High-throughput, robust glycomics profiling across many samples
Efficient comparison of glycosylation patterns across conditions, timepoints, or cohorts
Linkage-aware readouts when paired with chemical derivatization strategies
Considering MALDI–TOF–MS for a cohort or screening study? Tell us your sample type and question, we can suggest an analytical strategy.

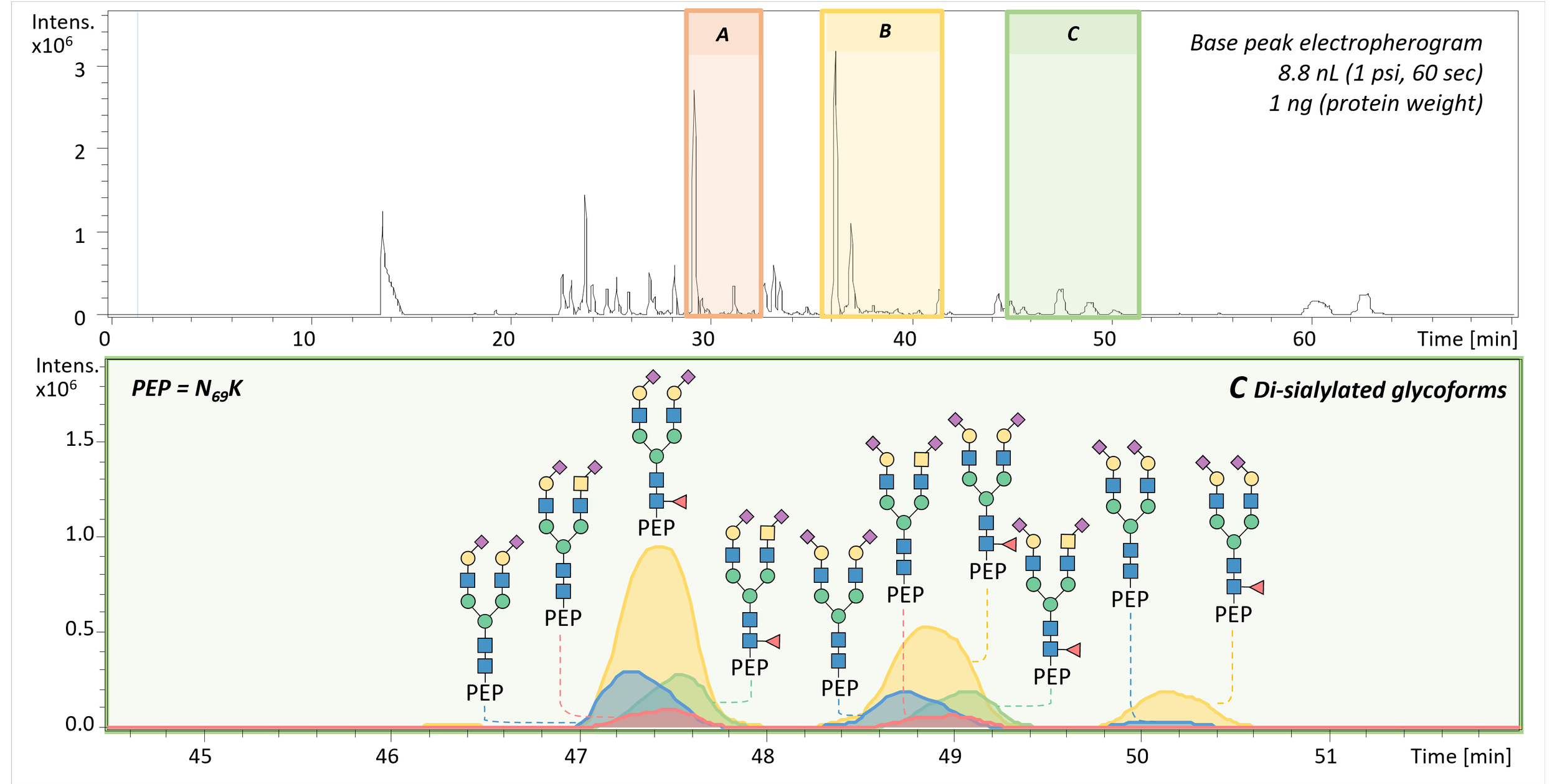
Isomer-resolved glyco(proteo)mics
In many studies, abundance changes are the first signal—glycans or glycopeptides becoming more or less prevalent across conditions. But for glycosylation, which structural variant changes can be just as important: different isomers and linkages can carry different biological implications, even when overall composition looks similar. Our workflows therefore aim to quantify abundance while adding isomer-level specificity where it strengthens interpretation.
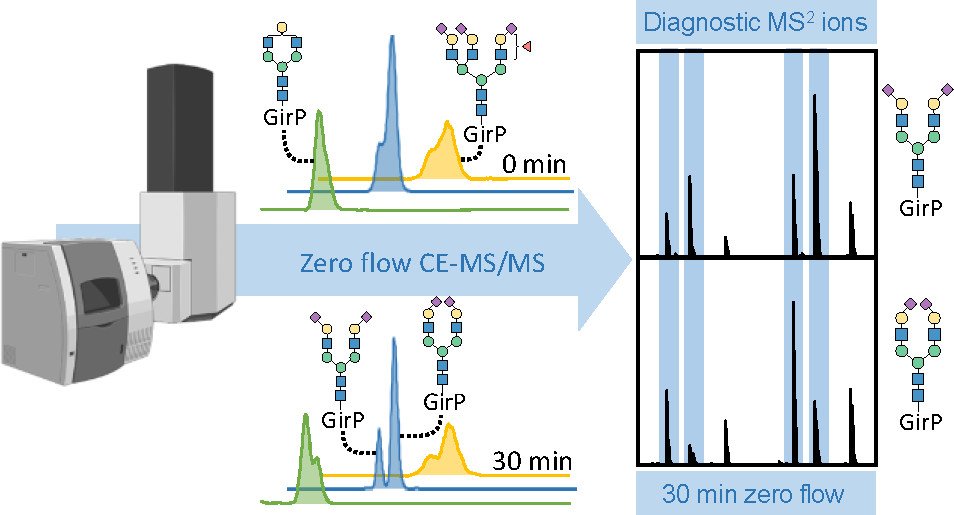
A central strength of our separation-based approach is that capillary electrophoresis (CE) can resolve isomeric species, including glycans and glycopeptides that differ only in subtle structural features. In particular, CE enables differentiation of sialic acid linkage isomers (α2,3 vs α2,6) with no additional sample preparation required, providing an orthogonal layer of information alongside abundance. When desired, we complement CE with chemical derivatization strategies that convert α2,3 and α2,6 sialylation into a diagnostic mass difference, enabling rapid discrimination by MS and supporting fast profiling by MALDI–TOF–MS. We also use advanced CE modes, including neutral, zero-flow CE–MS, to extend isomer separation further, supporting resolution of features such as arm occupancy and distinctions like core- vs antenna-fucosylation that can be critical for biological interpretation. These derivatization strategies can also translate to MALDI–MSI workflows, enabling spatial questions where linkage patterns matter in tissue context.
What this capability enables
Quantitative readouts that combine abundance with α2,6 vs α2,3 sialylation specificity
Deeper isomer separation with neutral/zero-flow CE–MS for additional structural distinctions
Derivatization-enabled fast profiling by MALDI–TOF–MS and compatibility with MALDI–MSI for spatial glycomics
Interested in adding isomer-level specificity to a glyco(proteo)mics question? Explore Applications for examples.
Spatial glycomics and tissue context (LCM + MSI)
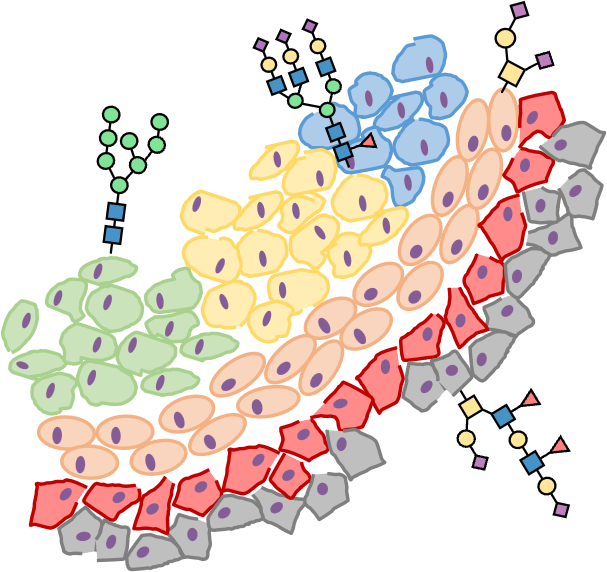
Spatial biology asks not only what changes, but where it changes. In the SUGAR Lab, we approach spatial glycosylation by combining laser capture microdissection (LCM) with sensitive downstream glycomics workflows, allowing us to interrogate tissue regions with higher specificity than bulk extraction. At present, we can perform N- and O-glycomics on microdissected material, linking glycan features to defined histological or microenvironmental regions and enabling studies of heterogeneity in disease tissue.
In parallel, the University Medical Center Groningen (UMCG) and Faculty of Science and Engineering (FSE) have invested in a state-of-the-art mass spectrometry imaging (MSI) platform to push spatial biology forward. The platform includes a Bruker timsTOF fleX MALDI-2 with TIMS, MALDI-2 post-ionization, and microgrid technology (enabling ~5 µm spatial resolution for imaging metabolites, drugs, lipids, and proteins), complemented by a MassTech atmospheric-pressure MALDI source compatible with the Orbitrap Exploris series for ultra-high mass resolution (reported up to 500,000) and ~10 µm spatial resolution. The added separation power of TIMS helps disentangle complex molecular signals and reduce interferences, while MALDI-2 boosts sensitivity, together creating a strong foundation for expanding spatial omics capability.
While we are not yet running full spatial glycomics on MSI as a routine workflow, establishing glycomics-capable MSI on this platform is a clear near-future direction, with the aim of bringing glycan biology into the same spatially resolved framework now transforming metabolite and lipid imaging.
What this capability enables
Region-specific N- and O-glycomics via laser capture microdissection
Tissue heterogeneity studies that connect glycosylation to histology and microenvironment
A pathway toward MSI-based spatial glycomics, leveraging high spatial resolution, TIMS separation, and MALDI-2 sensitivity
Interested in a tissue or spatial project? Explore Applications for examples or reach out via the project intake form to discuss feasiblity.
